Weave®, Spatial Insights at Enterprise Scale
Spatial biology is revolutionizing the drug discovery process across different therapeutic areas, such as oncology, immunology, and neurobiology, by adding cellular context to molecular omics layers.
Weave® is the industry-leading platform for spatial biology uniting people, data, and tools, by providing a collaborative and secure environment where massive multi-modal spatial datasets are turned into actionable insights.
Weave:
- Provides you with the tools to unlock the full potential of spatial biology.
- Multi-omic integration, multicellular neighborhood analysis, and AI/ML-native capabilities.
- Weave enables scalable analysis for any technology and multi-user teams.
- Enterprise-grade security: ISO 27001 and ISO 27018-certified.
- Trusted by top 20 Pharma and research teams worldwide
Key Takeaways
Any technology, any scale
Supports data from different spatial biology assays. Analyze single modalities or mix and match different spatial assays for comprehensive multi-omic characterization. Analyze hundreds of millions of cells across cohorts.
Multi-omics integration at full resolution
Integrate spatial multi-omics data to a common spatial footprint. All data presented at full resolution, with no image down sampling. State-of-the-art elastic registration algorithm that works across multiple images, vendors, and technologies.
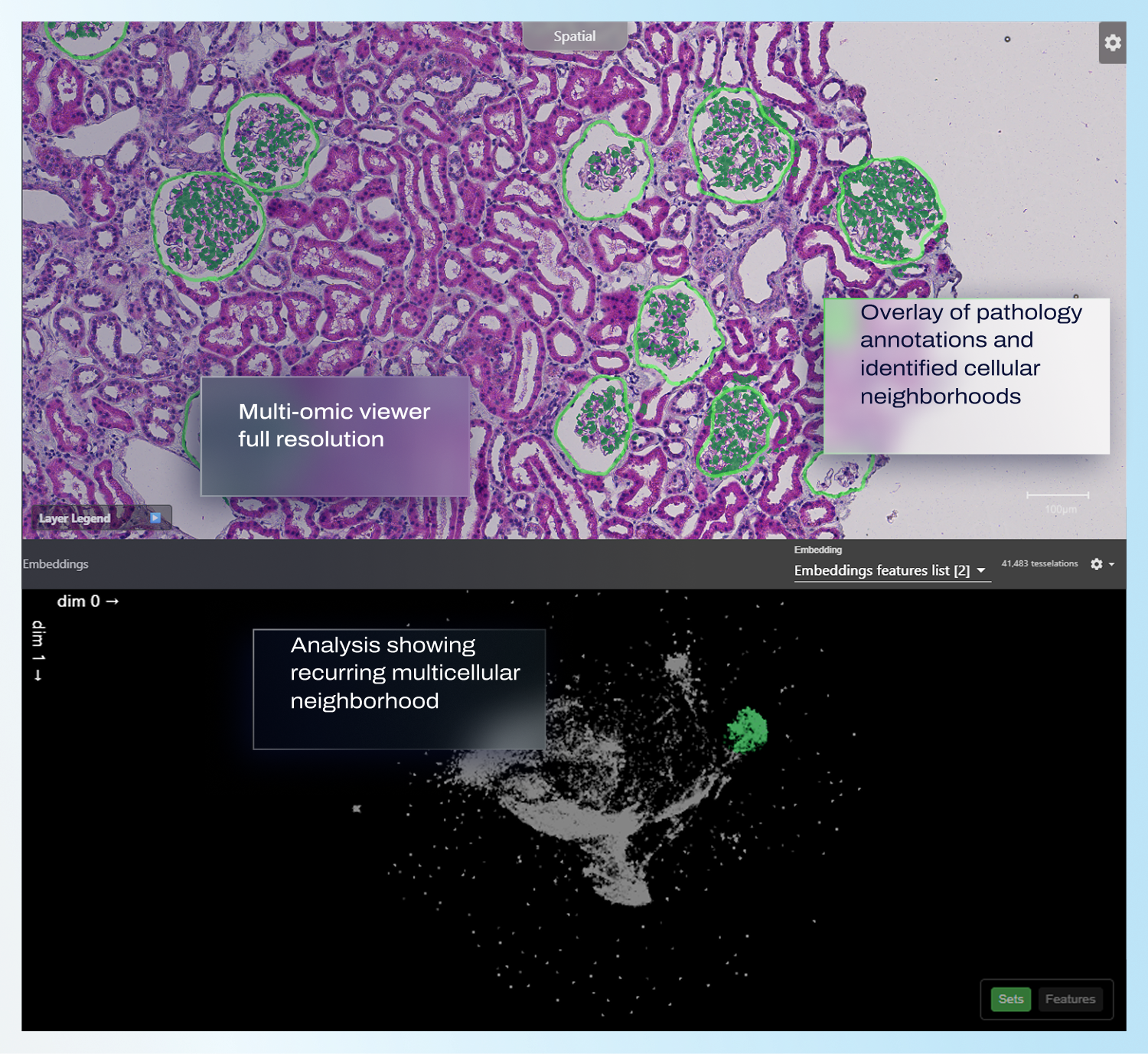
Multicellular neighborhood analysis
Proprietary algorithms to identify, analyze and interpret spatially recurring microenvironments across samples and cohorts. Identify cellular neighborhoods (also known as cell niches or spatial domains)associated with disease progression or pathology. Compare neighborhoods e.g. across responder vs non-responder cohorts with descriptive statistics.
Multi-user and multi-team
Collaborative, shareable, compliant workspaces: pathologists, biologists, computational biologists, and data leaders can collaborate in real-time on the same datasets. True democratization of spatial biology insights.
Built for the enterprise
ISO 27001 and ISO 27018-certified security and handling of PII for enterprise use. Single Sign-On integration following industry best practices. Scalable storage and computation resources. Hosted and on-premise deployment options. Natural language agent for guided data analysis and MCP server to integrate Weave in GenAI workflows.
Frequently Asked Questions
What spatial biology assays and technologies does Weave support?
Weave supports data from different commercially available spatial biology assays and vendors. You can analyze single modalities or mix and match different spatial assays for comprehensive multi-omic characterization: from spatial transcriptomics, proteomics, and metabolomics (mass spectrometry imaging-MSI), to histology, single-cell data, and clinical metadata.
A state-of-the-art elastic registration algorithm works across multiple images, vendors, and technologies.
Can Weave integrate with our existing pipelines and tools?
Yes. Weave is architected from the ground up to be tightly integrated with other systems, providing maximal flexibility through its own Python SDK. Connect to internal pipelines, catalogs, and compute environments via APIs and SDKs. Integrate results from in-house pipelines and third-party software, such as digital pathology tools (e.g., QuPath). Harmonized datasets and standardized workflows deploy back to your tools and data catalog. Insights compound across studies.
How does Weave handle large-scale, multi-team programs?
Whether analyzing a large cohort with one spatial assay or many different techniques on a smaller number of samples, Weave enables powerful and scalable analysis, efficient integration, and visualization, with team collaboration and data management wrapped in one package. FAIR data management lets you efficiently organize, search, and retrieve all the data and analysis results stored in Weave®. All datasets and downstream analysis results can be published to other teams within Weave following FAIR principles with ontology grounding.
For research use only. Not for diagnostic purposes. © 2026