Non-negative matrix factorization in MSI

In this blog post, we continue our overview of unsupervised data analysis approaches commonly used in MSI, diving deeper into factorization methods. This time we will take a closer look at the results of principal component analysis (PCA) in MSI, and compare these to the results of nonnegative matrix factorization (NMF).
This post is part of our series titled "Unsupervised learning on MSI data", which contains the following entries:
- Factorization of MSI data - part 1
- Non-negative matrix factorization in MSI (current post)
Table of contents
Introduction
In our last post on factorization, we explored a number of factorization methods commonly used in Mass Spectrometry Imaging (MSI) data analysis. This, of course, included the king of factorization in MSI, principal component analysis (PCA). As highlighted in our review paper, PCA is probably the most used unsupervised analysis technique in MSI, and is one of the most popular machine learning algorithms in general. PCA owes its royal status to a number of important strengths:
- it has a unique solution, meaning you get the same result everytime you run the algorithm (yes yes, except for component signs, but now you’re nitpicking, you can do a lot worse).
- it allows for a ranking of the “importance” of the components, importance here being the amount of variance a component explains in the data. A set of large peaks that changes throughout the tissue will most likely be of interest and will be ranked high in the results.
- if we are only interested in a limited number of components, e.g. the first 100 principal components, as is usually the case, these can be estimated very quickly even in large datasets.
- due to its relative simplicity, it is widely available in most data analysis packages.
There are some disadvantages, however, mainly in the interpretation of the results of a PCA analysis, especially in the context of MSI. Below, we recap the PCA results of the lymphoma tissue section that we introduced previously. Figure 1 shows the first 10 principal components, where the spatial expression images and their matching pseudo-spectra are shown on the left and right respectively. The pseudo-spectra show the biomelecular ions that are involved in the expression of the spatial pattern on the left. The way that these are often interpreted, is that the peaks in the pseudo-spectrum exhibit the spatial expression on the right. Things are usually more complex than that, however, as we’ll see below.
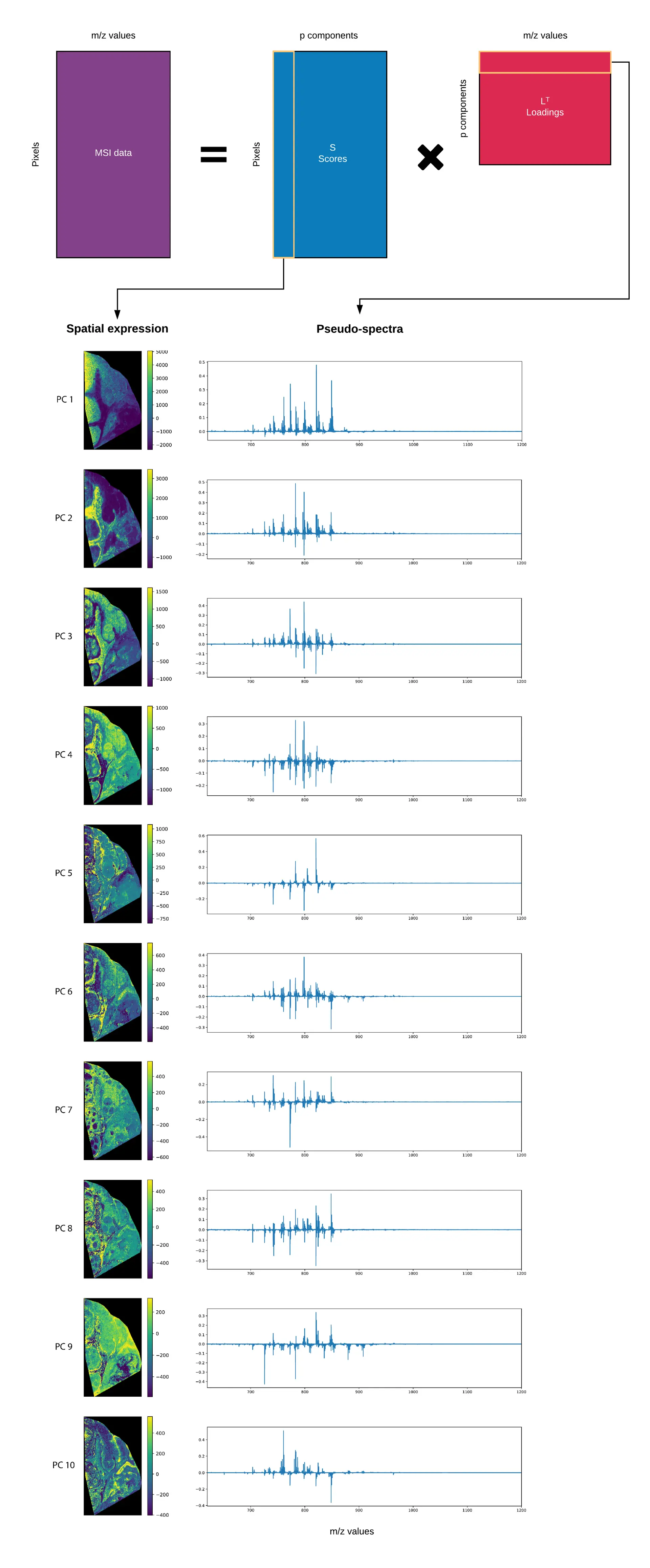
What's with all this negativity?
One feature of the PCA results that immediately sticks out when looking at the pseudo-spectra and spatial expression images, is the negative values that are present in the results. And this, while there are no negative values in the original MSI data, what is this!

But this is PCA and it doesn’t care about such trivial things as “physics” and “reality” – it is an abstract mathematical construct which captures as much variance per component as possible. PCA will do so by making linear combinations of the original variables, which together try to pack as much of the variability in the data as possible. These linear combinations can include negative coefficients, hence the negative values that we see in the results. Unfortunately, such negative coefficients have no straightforward physical interpretation in the case of MSI data.
Despite lacking a direct physical interpretation of coefficients, PCA will group together correlating variables, and thus there often is an underlying biochemical reason why variables are grouped together in a component. Furthermore, a negative coefficient could indicate a downregulation of a set of ions within the highly expressed region for the associated component as compared to the rest of the dataset. The main issues are (i) that this is not necessarily clear from the analysis and (ii) that PCA is not designed to accomodate such interpretation, especially for negative coefficients. What PCA will do, however, is point us to the variables, here the pixels and m/z bins, where interesting things are happening; i.e., PCA will uncover a linear subspace of interest in fancy terminology.
As we pointed out in our previous post, methods such as Varimax allow for a re-shuffling of the information in the components, such that the pseudo-spectra become less dense, and as such become easier to investigate. This still doesn’t not solve the problem of negative peaks though, and we don’t like all that negativity! Let’s put that frown upside down with some positivity, or, well, some non-negativity to be more exact. Zeros are ok, just no negative Nellies.
Positive thinking via non-negative matrix factorization
The problem of having undesirable negative values in a factorization result isn’t exclusive to MSI or mass spectrometry in general. Many other fields such as image and audio processing deal with data that is originally only positive in nature, and where it is desirable to have a factorization result that only contains positive values, aptly dubbed non-negative matrix factorization (NMF). Originally, the term positive matrix factorization had been coined, but as zeros do not identify as positive or negative, the more inclusive variant non-negative matrix factorization was subsequently adopted.
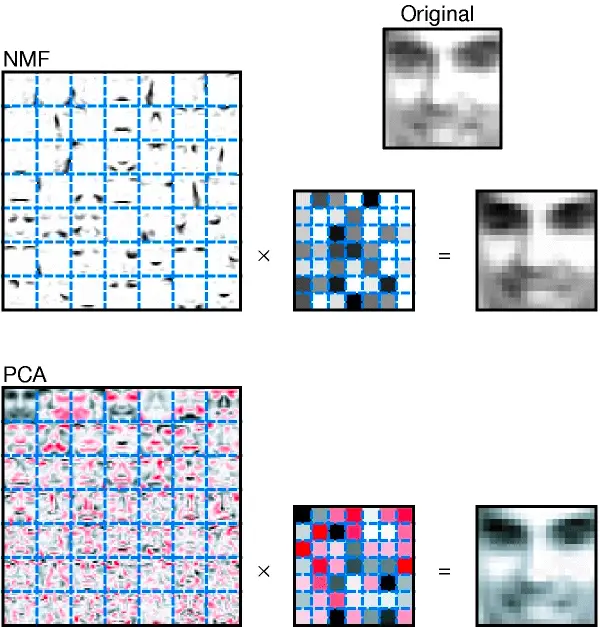
One of the hallmark papers for non-negative matrix factorization is that of Lee and Seung1, where both PCA and NMF are used for the decomposition of a large number of images of faces, with the goal of finding the underlying structure in these images.
The results are shown in Figure 3 above. Similar to the MSI results, the left image shows the spatial expression, the middle image is the equivalent of the pseudo-spectrum in MSI, and the right image shows a reconstructed image based on the components. In the PCA image, red pixels indicate negative coefficients.
In the results, NMF naturally splits up the spatial expressions into parts-based components for the eyes, nose, mouth, … (and also bunch of less-descriptive features). The original faces can then be recreated as additive combinations of these basis parts, making the reconstruction very easy to interpret. PCA, on the other hand, creates more holistic components that somewhat resemble face outlines (if you squint hard), with emphasis on combinations of various facial features (along with a large number of non-descriptive components). In comparison, these components are nowhere near as interpretable as those from NMF. We do note that the reconstructions from PCA are at least up to par with those of NMF, so it is mainly about how the information is broken down. In their work, Lee and Seung argue that the way NMF operates is also closer to how the human brain parses and represents information, as neurons cannot have negative firing rates. As such the brain likely operates in an additive way.
Non-negative matrix factorization is not a single technique like PCA, but rather is a group of algorithms that sets a goal i.e. to create a matrix factorization with only positive values and zeros, while trying to optimize a certain objective (function), and there are many different ways to do this. Moreover, it is possible to add additional constraints, such as sparsity into the objective function, which can further help in the interpretability of the results. For more information on this, we refer to our review paper on unsupervised analysis of MSI data. It is important to note, however, that for any of the different methods of calculating NMF, there is no unique, analytical solution like there is in PCA. These are all iterative approaches, and do not guarantee that the final solution is the optimal one. This also means that results may change from run to run, and will generally depend on the initial estimate from which the algorithm starts. This is in fact something where PCA (or usually its counterpart SVD) can help, namely to find good initial components (as we said before, PCA is really good at finding relevant variables), and then iteratively purify these to components that contain only zeros and positive values.
NMF applied in Mass Spectrometry Imaging
Below in Figure 4 we show 10 components of the NMF analysis on our lymphoma tissue. We use the scikit-learn implementation of NMF, calculating the first 100 components and using the ‘nndsvd’ (Nonnegative Double Singular Value Decomposition) as initialization, which promotes sparseness in the resulting components.
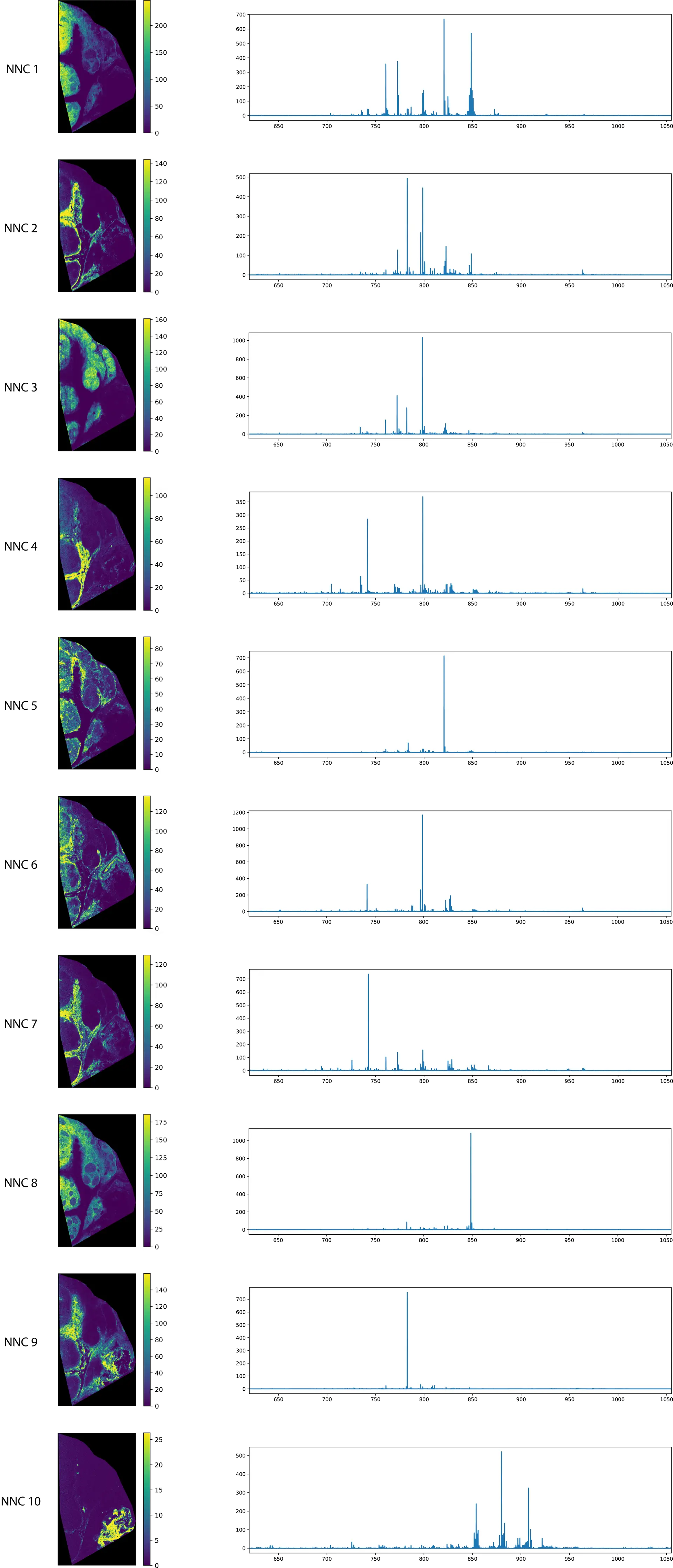
When inspecting these results, we immediately see that the pseudo-spectra look a lot less complex than those of PCA, and most importantly (and very unsurprisingly by this point), they are all positive, both in the expression image and the pseudo-spectra! Life goal achieved!

Similar to the results of the face decomposition, we see that the spatial expressions are much more localized to specific anatomical regions in the tissue, rather than having expressions throughout the whole tissue as we had in PCA. The pseudo-spectra are also sparser, containing a decidedly smaller number of peaks. Overall, we get a cleaner, more interpretable breakdown of our MSI data.
Despite the aforementioned boons, it is important to realize that peaks in NMF pseudo-spectra are not necessarily exclusive to their associated expression regions, nor should they follow the exact localization of the spatial expression images! We will illustrate this important and often misunderstood phenomenon in a subsequent post in this series, where we will talk about the infamous ghost peaks – spooky!
Conclusions
In this post we’ve gone deeper into how non-negative matrix factorization works, and how it can help in alleviating some of the interpretability issues of principal component analysis, by removing negative values from the spatial expression images and negative peaks from the pseudo-spectra, making the spectra more sparse.
If you are looking for some non-negative results from your MSI data analysis, make sure to contact us!